SOM cluster: 47
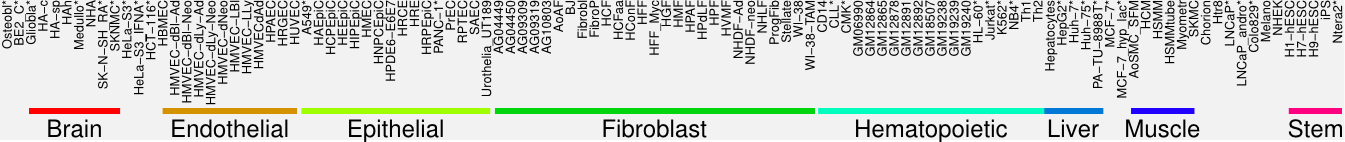
Cluster Hypersensitivity Profile
Genomic Location Trend
These plots show the distribution of the DHS sites surrounding the Transcript Start Site of the nearest gene.
Stats
Number of sites: 162Promoter: 71%
CpG-Island: 70%
Conserved: 70%
Enriched Motifs & Matches
Match Detail: [Jaspar]

| ||
|---|---|---|
| Sites: 84/100 | e-val: 6.7e-40 | ||
| Factor | e-val(match) | DB |
| SP1 | 0.00000000000093159 | JASPAR |
| INSM1 | 0.000061157 | JASPAR |
| Klf4 | 0.00012859 | JASPAR |
| Egr1 | 0.00026243 | JASPAR |
| CTCF | 0.002176 | JASPAR |

| ||
| Sites: 43/100 | e-val: 3.4e-36 | ||
| Factor | e-val(match) | DB |
| NFYA | 0.000000547 | JASPAR |
| NFIC | 0.0016919 | JASPAR |
| Pdx1 | 0.012749 | JASPAR |
| PBX1 | 0.029526 | JASPAR |
| GATA3 | 0.058936 | JASPAR |

| ||
| Sites: 14/100 | e-val: 1.8e-26 | ||
| Factor | e-val(match) | DB |
| znf143 | 0.000040204 | JASPAR |
| NFATC2 | 0.00046132 | JASPAR |
| FEV | 0.00080365 | JASPAR |
| Stat3 | 0.00096432 | JASPAR |
| SPI1 | 0.0069051 | JASPAR |

| ||
| Sites: 10/100 | e-val: 1.7e-19 | ||
| Factor | e-val(match) | DB |
| NFE2L2 | 0.00035168 | JASPAR |
| AP1 | 0.00035918 | JASPAR |
| Sox2 | 0.0033179 | JASPAR |
| Spz1 | 0.0085044 | JASPAR |
| NFIL3 | 0.015751 | JASPAR |

| ||
| Sites: 10/100 | e-val: 2.4e-18 | ||
| Factor | e-val(match) | DB |
| Myb | 0.000031885 | JASPAR |
| NFYA | 0.0021207 | JASPAR |
| Pdx1 | 0.0043652 | JASPAR |
| znf143 | 0.0072859 | JASPAR |
| Tcfcp2l1 | 0.0097899 | JASPAR |
BED file downloads
Top 10 Example Regions
| Location | Gene Link | Dist. |
|---|---|---|
| chr19: 21541660-21541810 | ZNF738 | 56.47 |
| chr1: 27019580-27019730 | PIGV | 57.14 |
| chr20: 32308020-32308170 | CBFA2T2 | 59.01 |
| chr20: 32308020-32308170 | PXMP4 | 59.01 |
| chr5: 139927100-139927250 | SNORA27 | 60.8 |
| chr19: 21688260-21688410 | ZNF429 | 61.48 |
| chr19: 12098320-12098470 | ZNF844 | 65.03 |
| chr19: 12098320-12098470 | ZNF433 | 65.03 |
| chr19: 12098320-12098470 | RNA5SP465 | 65.03 |
| chr19: 12098320-12098470 | CTD-2006C1.2 | 65.03 |