SOM cluster: 25
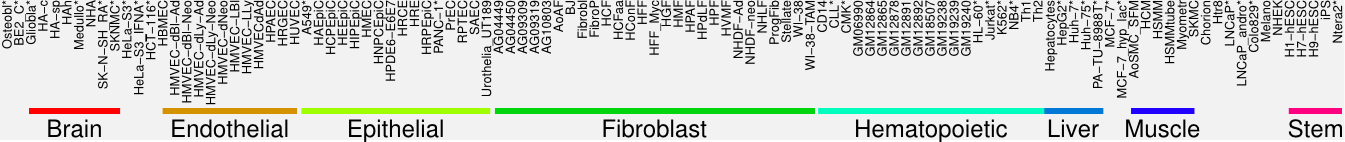
Cluster Hypersensitivity Profile
Genomic Location Trend
These plots show the distribution of the DHS sites surrounding the Transcript Start Site of the nearest gene.
Stats
Number of sites: 270Promoter: 33%
CpG-Island: 2%
Conserved: 66%
Enriched Motifs & Matches
Match Detail: [Jaspar]

| ||
|---|---|---|
| Sites: 89/100 | e-val: 0 | ||
| Factor | e-val(match) | DB |
| SPI1 | 0.0000000067926 | JASPAR |
| SPIB | 0.0000039257 | JASPAR |
| FEV | 0.0000043071 | JASPAR |
| ELF5 | 0.000065545 | JASPAR |
| ELK4 | 0.00013432 | JASPAR |

| ||
| Sites: 47/100 | e-val: 2.9e-32 | ||
| Factor | e-val(match) | DB |
| SP1 | 0.0000000012724 | JASPAR |
| RREB1 | 0.00000081686 | JASPAR |
| Pax4 | 0.0000051177 | JASPAR |
| Klf4 | 0.000012129 | JASPAR |
| PLAG1 | 0.00018734 | JASPAR |

| ||
| Sites: 28/100 | e-val: 0.00000029 | ||
| Factor | e-val(match) | DB |
| SPI1 | 0.00000085927 | JASPAR |
| ELK4 | 0.0000012654 | JASPAR |
| SPIB | 0.0000015457 | JASPAR |
| FEV | 0.0000027908 | JASPAR |
| Stat3 | 0.00002712 | JASPAR |

| ||
| Sites: 32/100 | e-val: 0.031 | ||
| Factor | e-val(match) | DB |
| RUNX1 | 0.00000000068778 | JASPAR |
| ZNF354C | 0.001335 | JASPAR |
| MYC::MAX | 0.011646 | JASPAR |
| RREB1 | 0.01782 | JASPAR |
| MAX | 0.019384 | JASPAR |

| ||
| Sites: 22/100 | e-val: 0.024 | ||
| Factor | e-val(match) | DB |
| PLAG1 | 0.0000010084 | JASPAR |
| INSM1 | 0.00012822 | JASPAR |
| Zfp423 | 0.0026381 | JASPAR |
| EBF1 | 0.0062849 | JASPAR |
| Klf4 | 0.0068238 | JASPAR |
BED file downloads
Top 10 Example Regions
| Location | Gene Link | Dist. |
|---|---|---|
| chr1: 147790800-147790950 | RP11-495P10.4 | 26.37 |
| chr1: 147790800-147790950 | RP11-495P10.5 | 26.37 |
| chr1: 156124180-156124330 | BGLAP | 46.02 |
| chr1: 156124180-156124330 | MEX3A | 46.02 |
| chr1: 156124180-156124330 | LMNA | 46.02 |
| chr1: 156124180-156124330 | SEMA4A | 46.02 |
| chrX: 129220900-129221050 | RAB33A | 46.46 |
| chrX: 129220900-129221050 | AIFM1 | 46.46 |
| chrX: 129220900-129221050 | BCORL1 | 46.46 |
| chrX: 129220900-129221050 | ELF4 | 46.46 |