SOM cluster: 2333
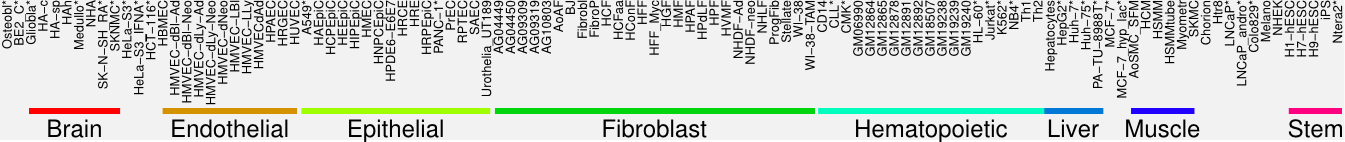
Cluster Hypersensitivity Profile
Genomic Location Trend
These plots show the distribution of the DHS sites surrounding the Transcript Start Site of the nearest gene.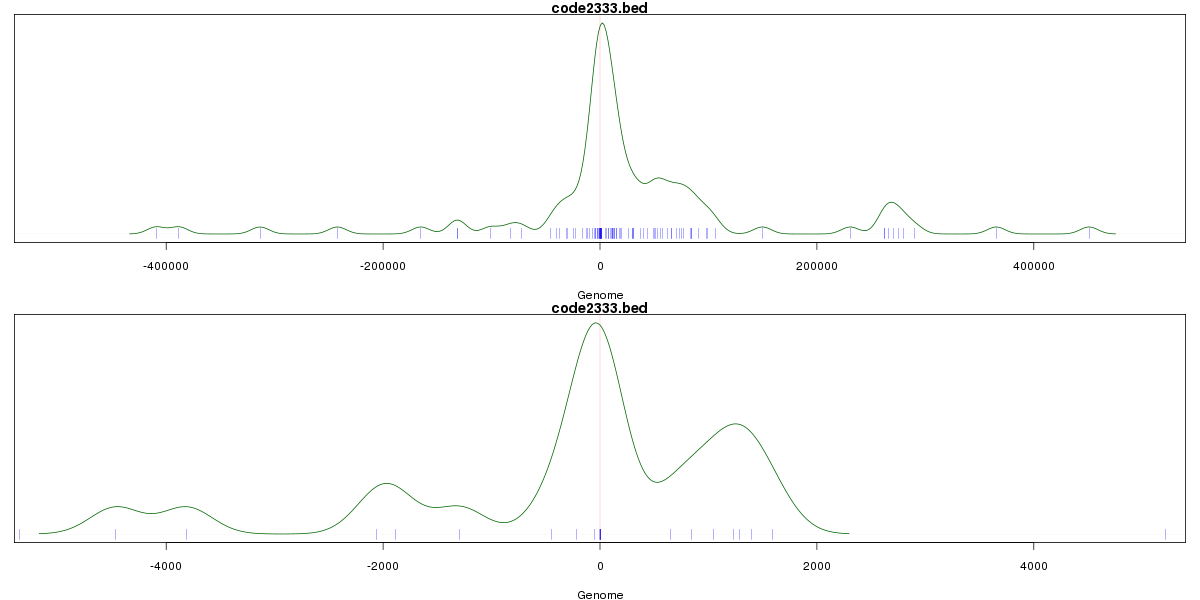
Stats
Number of sites: 316Promoter: 12%
CpG-Island: 5%
Conserved: 64%
Enriched Motifs & Matches
Match Detail: [Jaspar]
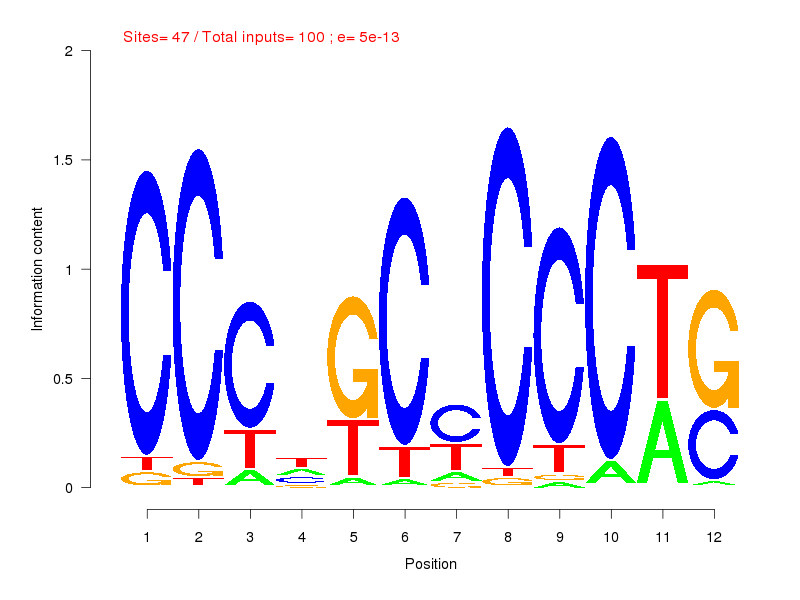
| ||
|---|---|---|
| Sites: 47/100 | e-val: 0.0000000000005 | ||
| Factor | e-val(match) | DB |
| SP1 | 0.0000000000053424 | JASPAR |
| INSM1 | 0.00000039554 | JASPAR |
| Tal1::Gata1 | 0.00014087 | JASPAR |
| CTCF | 0.00056917 | JASPAR |
| Klf4 | 0.0024918 | JASPAR |
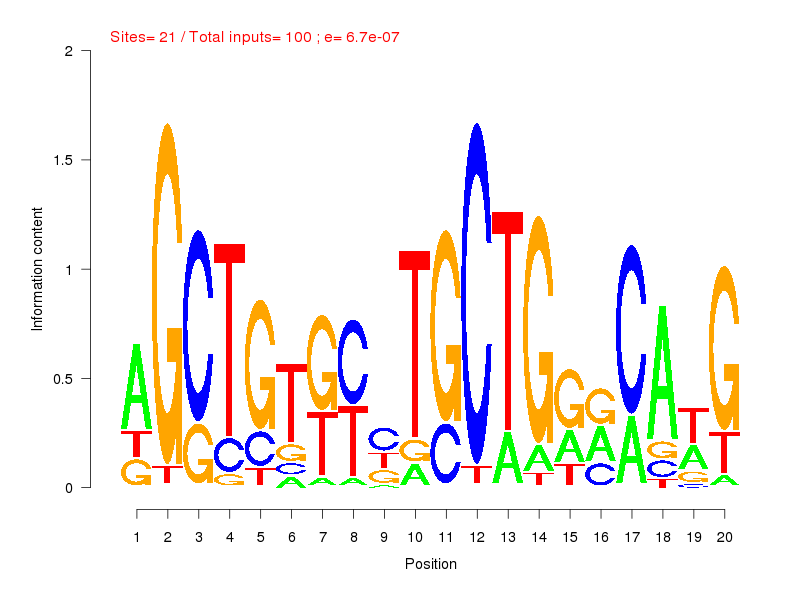
| ||
| Sites: 21/100 | e-val: 0.00000067 | ||
| Factor | e-val(match) | DB |
| Stat3 | 0.0015298 | JASPAR |
| znf143 | 0.01172 | JASPAR |
| NFATC2 | 0.023447 | JASPAR |
| FEV | 0.037657 | JASPAR |
| NFE2L2 | 0.039679 | JASPAR |
| ||
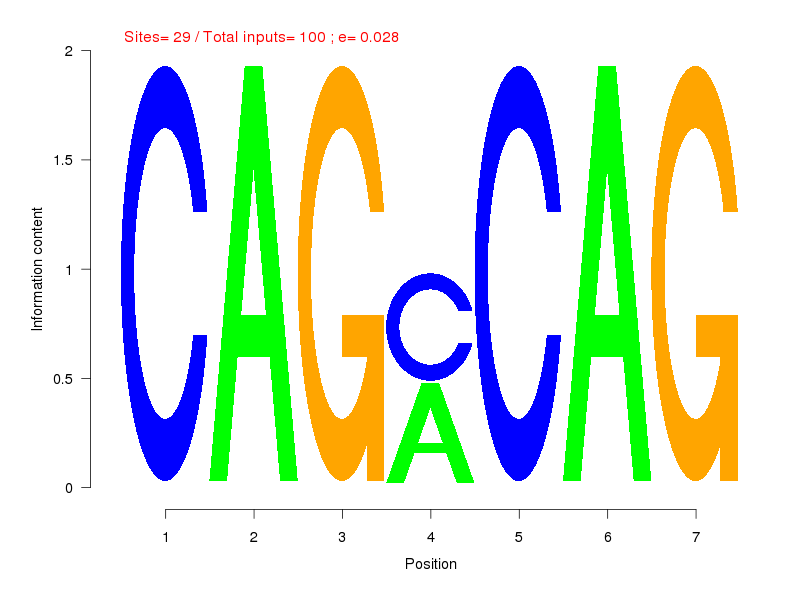
| Sites: 29/100 | e-val: 0.028 | ||
| Factor | e-val(match) | DB |
| Hand1::Tcfe2a | 0.0034515 | JASPAR |
| Tcfcp2l1 | 0.010183 | JASPAR |
| INSM1 | 0.016887 | JASPAR |
| NFIC | 0.020737 | JASPAR |
| REST | 0.033413 | JASPAR |
| ||
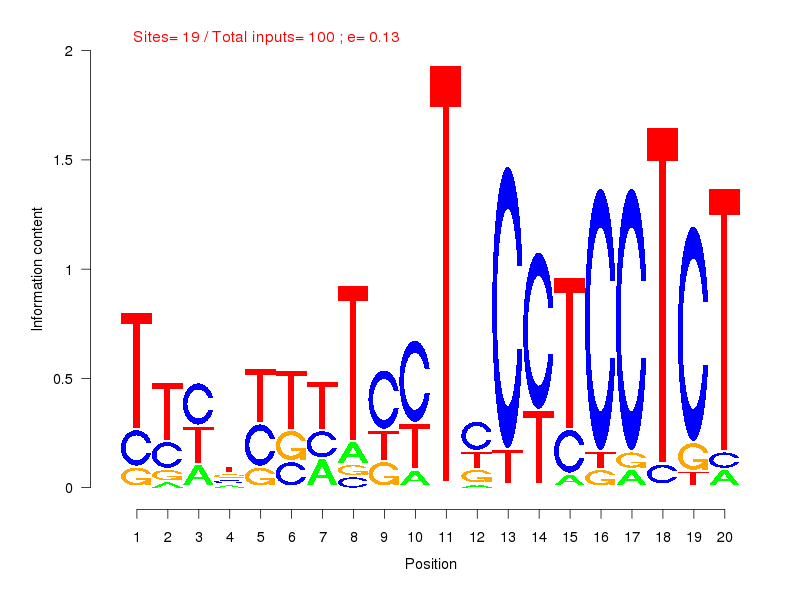
| Sites: 19/100 | e-val: 0.13 | ||
| Factor | e-val(match) | DB |
| EWSR1-FLI1 | 0.0000031496 | JASPAR |
| SPIB | 0.00027926 | JASPAR |
| Tal1::Gata1 | 0.0032852 | JASPAR |
| SPI1 | 0.0046002 | JASPAR |
| SP1 | 0.0050637 | JASPAR |
BED file downloads
Top 10 Example Regions
| Location | Gene Link | Dist. |
|---|---|---|
| chr1: 92311640-92311790 | TGFBR3 | 13.6 |
| chr1: 210405500-210405650 | SERTAD4-AS1 | 42.53 |
| chr1: 210405500-210405650 | HHAT | 42.53 |
| chr1: 210405500-210405650 | SERTAD4 | 42.53 |
| chr1: 210405500-210405650 | SYT14 | 42.53 |
| chr2: 47299145-47299295 | C2orf61 | 46.13 |
| chr11: 2019045-2019195 | TNNT3 | 47.56 |
| chr11: 2019045-2019195 | H19 | 47.56 |
| chr5: 178594505-178594655 | ADAMTS2 | 50.1 |
| chr5: 178594505-178594655 | ZNF354C | 50.1 |