SOM cluster: 2205
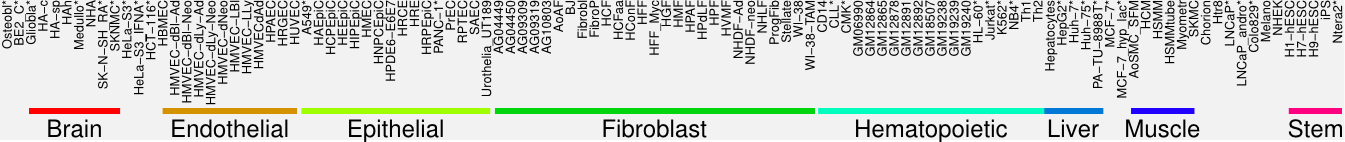
Cluster Hypersensitivity Profile
Genomic Location Trend
These plots show the distribution of the DHS sites surrounding the Transcript Start Site of the nearest gene.
Stats
Number of sites: 368Promoter: 5%
CpG-Island: 1%
Conserved: 57%
Enriched Motifs & Matches
Match Detail: [Jaspar]

| ||
|---|---|---|
| Sites: 47/100 | e-val: 0.000000059 | ||
| Factor | e-val(match) | DB |
| SOX10 | 0.0000024611 | JASPAR |
| SOX9 | 0.000047523 | JASPAR |
| Sox2 | 0.0016395 | JASPAR |
| NR4A2 | 0.0021238 | JASPAR |
| PPARG::RXRA | 0.0033637 | JASPAR |

| ||
| Sites: 11/100 | e-val: 0.00056 | ||
| Factor | e-val(match) | DB |
| Foxd3 | 0.000067 | JASPAR |
| IRF1 | 0.0010351 | JASPAR |
| FOXA1 | 0.0013855 | JASPAR |
| Foxq1 | 0.014321 | JASPAR |
| MEF2A | 0.014614 | JASPAR |
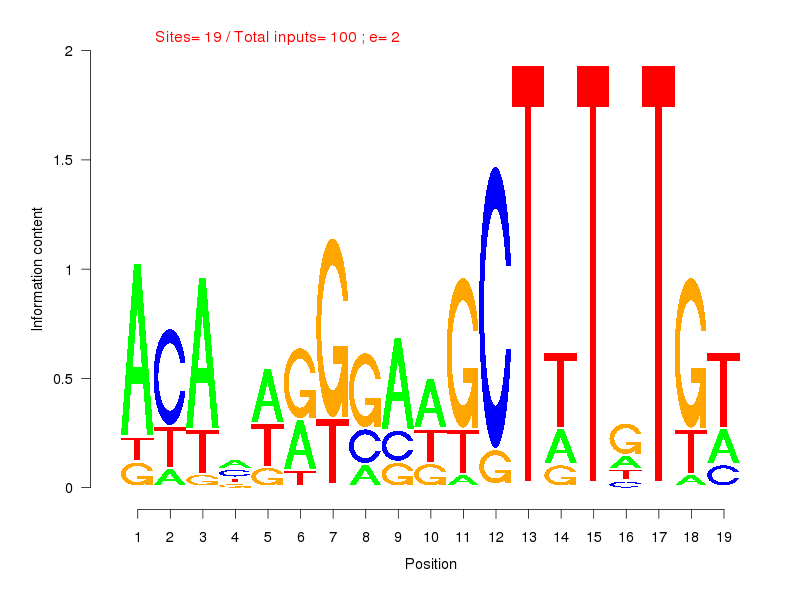
| ||
| Sites: 19/100 | e-val: 2 | ||
| Factor | e-val(match) | DB |
| SOX10 | 0.000029128 | JASPAR |
| Nr2e3 | 0.0003945 | JASPAR |
| SOX9 | 0.0026372 | JASPAR |
| Foxd3 | 0.0066525 | JASPAR |
| SPI1 | 0.0092577 | JASPAR |
BED file downloads
Top 10 Example Regions
| Location | Gene Link | Dist. |
|---|---|---|
| chr1: 203315640-203315790 | BTG2 | 26.09 |
| chr9: 116766545-116766695 | AMBP | 32.02 |
| chr4: 154170945-154171095 | TRIM2 | 49.26 |
| chr3: 37742380-37742530 | ITGA9 | 52.8 |
| chr3: 37742380-37742530 | AC093415.2 | 52.8 |
| chr6: 10375740-10375890 | TFAP2A-AS1 | 53.35 |
| chr6: 10375740-10375890 | TFAP2A | 53.35 |
| chr6: 10375740-10375890 | LINC00518 | 53.35 |
| chr12: 120658745-120658895 | GCN1L1 | 53.81 |
| chr1: 111005720-111005870 | PROK1 | 55.53 |