SOM cluster: 1625
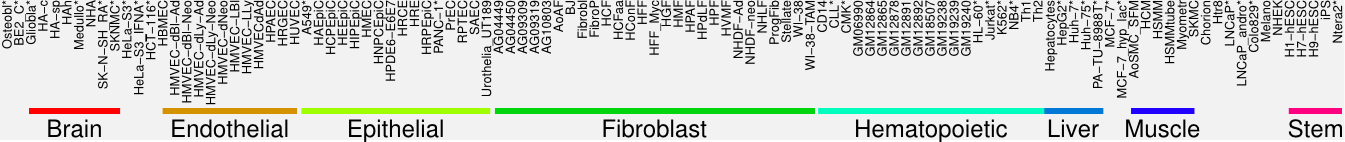
Cluster Hypersensitivity Profile
Genomic Location Trend
These plots show the distribution of the DHS sites surrounding the Transcript Start Site of the nearest gene.
Stats
Number of sites: 133Promoter: 1%
CpG-Island: 2%
Conserved: 47%
Enriched Motifs & Matches
Match Detail: [Jaspar]

| ||
|---|---|---|
| Sites: 47/100 | e-val: 6.7e-30 | ||
| Factor | e-val(match) | DB |
| AP1 | 0.0000000013888 | JASPAR |
| NFE2L2 | 0.000000002616 | JASPAR |
| NFE2L1::MafG | 0.000099138 | JASPAR |
| PBX1 | 0.0035981 | JASPAR |
| Pax2 | 0.011539 | JASPAR |

| ||
| Sites: 37/100 | e-val: 0.000029 | ||
| Factor | e-val(match) | DB |
| Foxa2 | 0.0000022604 | JASPAR |
| Foxq1 | 0.000092131 | JASPAR |
| FOXI1 | 0.0001421 | JASPAR |
| MEF2A | 0.0001513 | JASPAR |
| HNF1B | 0.00061171 | JASPAR |
BED file downloads
Top 10 Example Regions
| Location | Gene Link | Dist. |
|---|---|---|
| chr3: 158443640-158443790 | RARRES1 | 60.14 |
| chr3: 158443640-158443790 | MFSD1 | 60.14 |
| chr3: 158443640-158443790 | GFM1 | 60.14 |
| chr3: 158443640-158443790 | LXN | 60.14 |
| chr14: 53301300-53301450 | GNPNAT1 | 61.31 |
| chr14: 53301300-53301450 | FERMT2 | 61.31 |
| chr7: 100037360-100037510 | STAG3L5P | 61.41 |
| chr7: 100037360-100037510 | ZCWPW1 | 61.41 |
| chr7: 100037360-100037510 | PILRB | 61.41 |
| chr1: 172883760-172883910 | RP3-471M13.2 | 62.72 |