SOM cluster: 1369
Cluster Hypersensitivity Profile
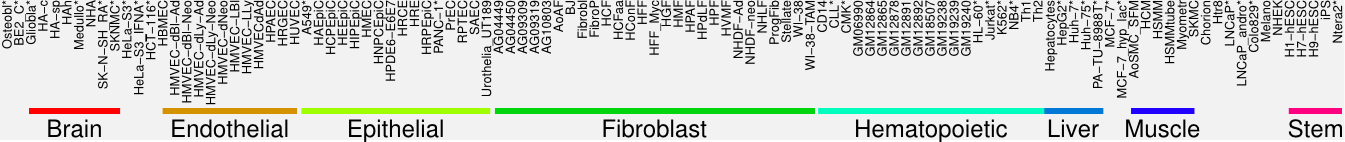
Genomic Location Trend
These plots show the distribution of the DHS sites surrounding the Transcript Start Site of the nearest gene.
Stats
Number of sites: 260Promoter: 0%
CpG-Island: 0%
Conserved: 48%
Enriched Motifs & Matches
Match Detail: [Jaspar]

| ||
|---|---|---|
| Sites: 83/100 | e-val: 0 | ||
| Factor | e-val(match) | DB |
| Myf | 0.0000000001583 | JASPAR |
| NHLH1 | 0.00000066821 | JASPAR |
| TAL1::TCF3 | 0.0016939 | JASPAR |
| Myb | 0.0058928 | JASPAR |
| CTCF | 0.0103 | JASPAR |

| ||
| Sites: 58/100 | e-val: 0.00000000054 | ||
| Factor | e-val(match) | DB |
| Zfx | 0.0003163 | JASPAR |
| Stat3 | 0.0016214 | JASPAR |
| INSM1 | 0.002499 | JASPAR |
| Klf4 | 0.0031484 | JASPAR |
| Hand1::Tcfe2a | 0.003963 | JASPAR |

| ||
| Sites: 29/100 | e-val: 0.027 | ||
| Factor | e-val(match) | DB |
| TLX1::NFIC | 0.00000000068231 | JASPAR |
| INSM1 | 0.00000021358 | JASPAR |
| ESR1 | 0.00068371 | JASPAR |
| PPARG::RXRA | 0.0008746 | JASPAR |
| ESR2 | 0.0021384 | JASPAR |

| ||
| Sites: 21/100 | e-val: 0.042 | ||
| Factor | e-val(match) | DB |
| SP1 | 0.026992 | JASPAR |
| Myf | 0.039595 | JASPAR |
| INSM1 | 0.05526 | JASPAR |
| NFYA | 0.057259 | JASPAR |
| SPIB | 0.0721 | JASPAR |
BED file downloads
Top 10 Example Regions
| Location | Gene Link | Dist. |
|---|---|---|
| chr11: 66719405-66719555 | SYT12 | 39.09 |
| chr1: 203021000-203021150 | PPFIA4 | 46.37 |
| chr1: 203021000-203021150 | ADORA1 | 46.37 |
| chr1: 203021000-203021150 | RP11-335O13.8 | 46.37 |
| chr1: 203021000-203021150 | MYOG | 46.37 |
| chr2: 29621780-29621930 | ALK | 49.3 |
| chr17: 76447180-76447330 | DNAH17 | 53.64 |
| chr2: 208116440-208116590 | AC007879.2 | 57.06 |
| chr2: 208116440-208116590 | AC007879.3 | 57.06 |
| chr2: 208116440-208116590 | AC007879.5 | 57.06 |